Nieuwe sluiproute door het kernporiecomplex ontdekt
Onderzoekers van de Faculteit Wiskunde en Natuurwetenschappen van de Rijksuniversiteit Groningen en van het Universitair Medisch Centrum Groningen hebben een nieuwe sluiproute blootgelegd waarmee membraaneiwitten naar de binnenste kernmembraan worden geloodst. Anders dan bij de bekende methode blijkt de ‘adrescode’ van het eiwit bevestigd aan een lange sliert, die zich door de membraanporie heen ’wiebelt’. Deze ontdekking wordt express-on-line gepubliceerd door het wetenschappelijke tijdschrift Science op 9 juni om 21.00 uur.
De Groningse onderzoekers hebben bestudeerd hoe bepaalde membraaneiwitten hun weg vinden naar de juist plek in een cel. De celkern en de kernporiecomplexen zijn aanwezig in alle cellen van plant, dier en mens en de functie ervan is cruciaal voor het leven. Bij disfunctioneren sterft de cel. Fundamenteel onderzoek als dit is een investering in de toekomst en heeft in dit specifieke geval waarschijnlijk toepassingsmogelijkheden in de geneeskunde.
Adrescode
Being in the right place at the right time is voor een eiwit van groot belang. De timing en locatie zijn essentieel voor het kunnen uitvoeren van zijn taak. Dit is geen nieuw inzicht, en van veel eiwitten is reeds bekend hoe en wanneer ze hun juiste locatie bereiken. Vaak is er een signaal gecodeerd op het oppervlak van het eiwit. Specifieke transporteiwitten herkennen deze ‘adrescode’ en kunnen het eiwit vervolgens op de juiste plaats in de cel brengen.
‘Linker’
De locatie waar de onderzoekers in geïnteresseerd waren, is het membraan aan de binnenkant van de celkern, de kern waar ook het DNA ligt. Om daar te komen moeten eiwitten via transport kanaaltjes, de kernporiën, naar binnen worden gebracht. Wat de Groningse onderzoekers nu hebben gevonden, is een adrescode die geheel anders is dan de al bekende adrescodes voor transport door de kernporiën. De nieuw ontdekte adrescode blijkt te hangen aan een lange zogeheten ‘linker’. Samenwerking tussen de drie eerste auteurs, Anne Meinema, Justyna Laba en Astri Hapsari was de sleutel tot het succes. De code is specifiek voor een aantal eiwitten dat in de membraan rondom de kern van een cel zit.
Sluiproute
De lange en flexibele linker kan gemakkelijk uitstrekken en opvouwen en kan hierdoor de adrescode een eind bij de membranen vandaan brengen. De linker ‘wiebelt’ zich een weg door het kernporie complex, zodat de adrescode, met daaraan de eiwitten die het transport verzorgen, op een efficiënte manier door het kanaal kunnen gaan. Daarmee is een nieuwe sluiproute door het kernporiecomplex ontdekt.
Bij de illustratie
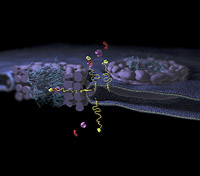
Te zien zijn twee kernporiecomplexen (paars met blauw) in de membraan rondom de kern en de route die een membraaneiwit (geel) volgt om aan de binnenkant van de kern te komen. Aangegeven is hoe transporteiwitten (rood en paars) de adrescode herkennen aan het einde van de lange linker. De linker wiebelt zich een weg door het kernporie-complex, zodanig dat de transporteiwitten door de binnenkant van het kanaal kunnen gaan.
Noot voor de redactie
• Nadere informatie: Dr. Liesbeth Veenhoff, tel. 050- 363 4187; l.m.veenhoff rug.nl
• Publicatie: Long Unfolded Linkers Facilitate Membrane Protein Import Through the Nuclear Pore Complex in Science 9 juni 2011;
• Auteurs: Anne C. Meinema, Justyna K. Laba, Rizqiya A. Hapsari, Renee Otten, Frans A. A. Mulder, Annemarie Kralt, Geert van den Bogaart, C. Patrick Lusk, Bert Poolman, Liesbeth M. Veenhoff
• Betrokken afdelingen: Biochemistry and Biophysical Chemistry, RUG; Neuroscience, Universitair Medisch Centrum Groningen; Cell Biology, Yale School of Medicine, VS.
Meer nieuws
-
08 juni 2026
Duurzaam samenleven op de savanne
-
02 juni 2026
Minuscuul reukspoortje ontrafeld
