Onderzoekers identificeren doelwitten voor geneesmiddelen tegen coronavirus

Wetenschappers van de RUG hebben, samen met collega’s van het IIMCB in Warschau (Polen) en Leiden UMC, mogelijke aangrijpingspunten gevonden voor geneesmiddelen in het RNA van het SARS-CoV2 coronavirus dat verantwoordelijk is voor de pandemie. Hun resultaten, die nog moeten worden beoordeeld door collega’s, laten zien welk deel van het genetisch materiaal van het virus een aantrekkelijk doelwit is voor bestrijding van het virus. De informatie kan helpen bij onderzoek naar een geneesmiddel voor het coronavirus.
Het manuscript met de onderzoeksresultaten is op maandag 15 juni geplaatst op de bioRxiv preprint server. Wetenschappers gebruiken dit soort preprint-servers om collega’s snel toegang te geven tot hun werk. Dat is belangrijk in onderzoek naar het coronavirus, aangezien de gebruikelijke procedure om wetenschappelijke resultaten te publiceren maanden kan duren. Maar omdat het manuscript nog niet is beoordeeld door experts kan het zijn dat er nog correcties of verbeteringen in de tekst worden gemaakt. De onderzoekers hebben hun werk inmiddels ingediend bij een tijdschrift dat de beoordeling zal uitvoeren. Momenteel wordt nieuwe informatie over het coronavirus al snel door de media opgepikt, ook als het om preprints gaat.
Het onderzoek werd geleid door Danny Incarnato van de RUG. Zijn lab onderzoekt de structuur van RNA, het genetisch materiaal van virussen zoals SARS-CoV2. Dit RNA bevat de genetische code voor het maken van eiwitten. Maar daarnaast bevatten de RNA-moleculen nog een tweede laag van informatie: delen van het enkelstrengs RNA die deze code bevat kunnen op zichzelf terugvouwen wat complexe 2D- of 3D-structuren oplevert. ‘Deze structuren zijn belangrijk voor allerlei aspecten van de levenscyclus van het virus, zoals de verspreiding en besmettelijkheid’, legt Incarnato uit.
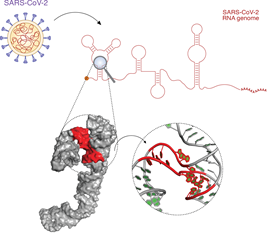
Hij gebruikte een bekende techniek om vast te stellen welke delen van het RNA gevouwen zijn. Collega’s van het International Institute of Molecular and Cell Biology (IIMCB) in Warschau zijn experts in het voorspellen van de 3D- structuren die de gevouwen delen vormen. ‘Dat liet zien waar er holtes aanwezig zijn in de structuur, waarin kleine moleculen zich kunnen binden’, vertelt Incarnato. Het binden van een klein molecuul in zo’n holte kan de werking van het RNA verstoren en daardoor ook het virus. ‘We weten niet in welke holtes dat kan gebeuren, maar door ze allemaal in kaart te brengen krijgen ontwikkelaars van geneesmiddelen een lijst met doelwitten die ze kunnen onderzoeken.’
De onderzoekers keken speciaal naar structuren die aanwezig zijn in verschillende typen coronavirussen: structuren die overal aanwezig zijn zullen vermoedelijk minder gemakkelijk veranderen door mutaties. Dit is belangrijk, aangezien RNA virussen snel muteren en zo ongevoelig kunnen worden voor geneesmiddelen of vaccins. ‘Een aantal van de 3D-structuren die we hebben gezien lijken nauwelijks te veranderen, zelfs niet als er nieuwe mutaties in het RNA ontstaan’, legt Incarnato uit. Dat is een groot voordeel van het gebruik van middelen gericht tegen deze 3D RNA-structuren bij de bestrijding van een virus.
Maar er zijn ook delen van het RNA niet gevouwen. Die gebieden zijn een mogelijk doelwit voor zogeheten antisense oligonucleotide behandeling. Die bestaat uit een klein kunstmatig DNA-molecuul dat zich bindt aan het virus-RNA. De dubbelstrengs structuur die dan ontstaat is gevoelig voor afbraak door enzymen die van nature voorkomen in onze cellen. Incarnato: ‘De afgelopen paar jaar is er een aantal van dit soort antisense RNA behandelingen goedgekeurd, het is een veelbelovende aanpak.’
Het onderzoek bestond uit een combinatie van laboratoriumwerk (het isoleren van RNA en het vinden van de gevouwen delen) en computerwerk (het uitrekenen van de 3D structuur en identificeren van de holtes). De resultaten kunnen het onderzoek naar nieuwe geneesmiddelen tegen het pandemische virus versnellen. Incarnato en zijn collega’s zijn zelf al op zoek naar zulke geneesmiddelen. Hij is er van overtuigd dat de structuren die in het artikel beschreven zijn kloppen. ‘We hebben ze vergeleken met bekende structuren van verschillende coronavirussen en we vonden een goede overeenkomst.’ Maar, besluit hij, het manuscript op de preprint-server is werk in uitvoering: de collegiale beoordeling kan onderdelen ervan veranderen. ‘Dat is nu eenmaal hoe wetenschap werkt: je hebt collega’s nodig die details zien waar je zelf overheen hebt gekeken. Ik ben er van overtuigd dat dit soort controle essentieel is.’
Meer nieuws
-
01 juni 2026
Materiaal met eigenschappen bepaald door structuur
