Development of Coarse-Grain Models
To extend the accessible time- and length scales of molecular simulations, the Berendsen Center actively develops coarse-grain (CG) models. In CG models, groups of atoms are united into CG beads with effective interactions betwee them. The interactions can be calibrated based on higher resolution models (bottom-up) or using experimental data (top-down), or a combination of both. When done carefully, CG models allow for a speed up of several orders of magnitude while keeping sight of the atomistic details.
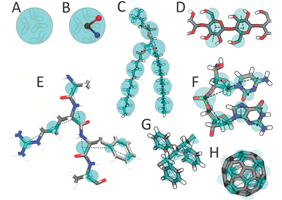
In the Molecular Dynamics group, the Martini model has been developed. The Martini forcefield is a CG force field suited for molecular dynamics simulations of biomolecular systems. The force field has been parametrized in a systematic way, combining top-down and bottum-up strategies: Non-bonded interactions are based on the reproduction of experimental partitioning free energies between polar and apolar phases of a large number of chemical compounds, whereas bonded interactions are derived from reference all-atom simulations.
The Martini model uses a four-to-one mapping, i.e. on average four heavy atoms and associated hydrogens are represented by a single interaction center. In order to keep the model simple, only four main types of interaction sites are defined: polar, non-polar, apolar, and charged. Each particle type has a number of subtypes, which allow for an accurate representation of the chemical nature of the underlying atomistic structure.
Due to its simplicity and transferability, the Martini force field is globally used and applied to a wide range of systems, from proteins to lipids and from polymers to nanoparticles. For more details, please visit the Martini website.
Key publications:
- S.J. Marrink, H.J. Risselada, S. Yefimov, D.P. Tieleman, A.H. de Vries. The MARTINI forcefield: coarse grained model for biomolecular simulations. JPC-B, 111:7812-7824, 2007. abstract
- L. Monticelli, S.K. Kandasamy, X. Periole, R.G. Larson, D.P. Tieleman, S.J. Marrink. The MARTINI coarse grained forcefield: extension to proteins. JCTC, 4:819-834, 2008. abstract
- C.A. Lopez, A. Rzepiela, A.H. de Vries, L. Dijkhuizen, P.H. Huenenberger, S.J. Marrink. The Martini coarse grained force field: extension to carbohydrates. J. Chem. Th. Comp., 5:3195-3210, 2009. abstract
- S.O. Yesylevskyy, L.V. Sch ä fer, D. Sengupta, S.J. Marrink. Polarizable water model for the coarse-grained Martini force field. PLoS Comp. Biol, 6:e1000810, 2010. open access
- D.H. de Jong, G. Singh, W.F.D. Bennett, C. Arnarez, T.A. Wassenaar, L.V. Schäfer, X. Periole, D.P. Tieleman, S.J. Marrink. Improved parameters for the Martini coarse-grained protein force field, JCTC, 9:687–697, 2013. open access
- S.J. Marrink, D.P. Tieleman. Perspective on the Martini model. Chem. Soc. Rev., 42:6801-6822, 2013. open access
- C. Arnarez, J.J. Uusitalo, M.F. Masman, H.I. Ingólfsson, D.H. de Jong, M.N. Melo, X. Periole, A.H. De Vries, S.J. Marrink. Dry Martini, a coarse-grained force field for lipid membrane simulations with implicit solvent. JCTC, 11:260–275, 2015. abstract
- J.J. Uusitalo, H.I. Ingólfsson, P. Akhshi, D.P. Tieleman, S.J. Marrink. Martini coarse-grained force field: extension to DNA. JCTC 11:3932-3945, 2015. open access
In the Molecular Mechanics group, further speed up is obtained by a supra-CG model with implicit solvent.
Key publications:
- Ghavami, A., van der Giessen, E., & Onck, P. R. (2013). Coarse-Grained Potentials for Local Interactions in Unfolded Proteins. Journal of Chemical Theory and Computation, 9(1), 432-440. 10.1021/ct300684j
- Ghavami, A., Veenhoff, L. M., van der Giessen, E., & Onck, P. R. (2014). Probing the Disordered Domain of the Nuclear Pore Complex through Coarse-Grained Molecular Dynamics Simulations. Biophysical Journal, 107(6), 1393-1402. 10.1016/j.bpj.2014.07.060
